Article
Special Issues
Spectroscopy Supplements
Advanced Antibody–Drug Conjugate Structural Characterization by Sheathless Capillary Electrophoresis–Tandem Mass Spectrometry Using Complementary Approaches
Author(s):
Antibody drug conjugates (ADCs) are an emerging category of biotherapeutic products based on monoclonal antibodies (mAbs) coupled to powerful cytotoxic drugs. The production of ADCs entails the formation of species with different number of conjugates drugs. The heterogeneity of ADCs species add to the complexity originating from the mAbs microvariability. Sheathless capillary electrophoresis-mass spectrometry (sheathless CE-MS) using complementary approaches was used to perform a detail characterization of brentuximab vedotin (Adcetris, Seattle Genetics). Sheathless CE-MS instrument used as nanoESI infusion platform was involved to perform the intact and middle-up analysis in native MS conditions. The nanoESI infusion approaches enabled estimation of the average drug to antibody ratio (DAR) alongside to drug load distribution. Sheathless CZE-MS/MS method developed was used to obtain from a single injection the characterization of the amino acid sequence with complete sequence coverage. In addition glycosylation and drug-loaded peptides could be identified from MS/MS spectra revealing robust information regarding their localizations and abundances. Drug-loaded peptide fragmentation mass spectra study demonstrated drug-specific fragments reinforcing the identifications confidence. Results reveal the ability of sheathless CZE-MS/MS method to characterize ADCs primary structure in a single experiment.
Antibody–drug conjugates (ADCs) are an emerging category of biotherapeutic products based on monoclonal antibodies (mAbs) coupled to powerful cytotoxic drugs. The production of ADCs entails the formation of species with different number of conjugates drugs. The heterogeneity of ADC species add to the complexity originating from the mAb microvariability. Sheathless capillary electrophoresis–mass spectrometry (sheathless CE–MS) using complementary approaches was used to perform a detailed characterization of brentuximab vedotin. A sheathless CE–MS instrument used as a nanoelectrospray ionization (nanoESI) infusion platform was involved to perform the intact and middle-up analysis in native mass spectrometry (MS) conditions. The nanoESI infusion approaches enabled estimation of the average drug-to-antibody ratio (DAR) and the drug load distribution. A single injection with a sheathless capillary zone electrophoresis (CZE)–MS/MS method was sufficient to characterize the amino acid sequence with complete sequence coverage. In addition glycosylation and drug-loaded peptides could be identified from MS/MS spectra, which revealed robust information about their localizations and abundances. A drug-loaded peptide fragmentation mass spectra study yielded drug-specific fragments, which reinforced the confidence about the identifications. The results reveal the ability of the sheathless CZE–MS/MS method to characterize an ADC’s primary structure in a single experiment.
The recent approval by the Food and Drug Administration (FDA) of brentuximab vedotin (Adcentris, Amgen) and trastuzumab emtansine (Kadcyla, Genentech/Roche) demonstrated the potential of antibody-drug conjugates (ADCs) as the next-generation of biotherapeutic products (1). Currently more than 50 ADCs are in clinical trials, which suggests a wider application of the biopharmaceutical industry in the near future (2). ADCs are based on monoclonal antibodies (mAbs) conjugated to highly potent cytotoxic drugs, mainly through covalent coupling. This format combines the extreme potency of the conjugated drug with the specificity for the targeted antigen provided by the mAbs (3).
The coupling of the drug through chemical reaction leads to the production of different species that correspond to the same ADC with a different number of conjugated drugs. In addition for a given species, the same number of drug molecules can be linked to different residues on the peptide backbone. On a macroscale level, this heterogeneity is characterized by different parameters like the drug–antibody ratio (DAR), which are considered as critical quality attributes (CQA) in the case of ADCs (4). It is important to take into account this variability is contributing to the microheterogeneities that naturally originate from the use of the mAbs format, such as glycosylation and other types of post-translational modifications (PTMs) (5). Various analytical techniques are required for comprehensive characterization of an ADC’s primary structure, which can require a good deal of both time and sample, especially during early stage development. Mass spectrometry (MS) is a versatile and efficient tool to study the structure of protein and characterize modifications (6–8). The technique yields precious information regarding the structure of proteins as well an exceptional selectivity. Therefore, it is extensively used to characterize therapeutic proteins in combination with a separation technique (9). Our group has developed a method based on sheathless capillary zone electrophoresis coupled to tandem mass spectrometry (CZE–MS/MS) for the comprehensive characterization of the primary structure of mAbs. This method proved to be capable, in a single injection, of characterizing the amino acid sequence of mAbs with complete sequence coverage and also of obtaining the glycosylation profile to study selected PTM hotspots (10). This analytical workflow could be further adapted to the primary structure assessment of biosimilars in comparison to their innovator mAbs (11).
Recently our group developed a sheathless CE–MS/MS analytical workflow, adapted to the additional complexity induced by the analysis of ADCs, to provide in a single analysis the characterization of this type of therapeutic protein (12). This analytical method, based on a derived bottom-up proteomic workflow, is designed to provide detailed information about the location on the peptide backbone of the conjugated drugs and a site-specific estimation of the conjugation reaction yield. The ADC we considered is brentuximab vedotin (BV), which was approved in 2011 by the FDA and successively by the European Medicine Agency (EMA) for the treatment of Hodgkin’s lymphoma and systemic anaplastic large-cell lymphoma, which represents a conceptual model for ADCs approved in the future. BV is composed of the tubulin inhibitor monomethyl auristatin E (MMAE) conjugated to the chimeric mAb cAC10 via a protease-cleavable maleimidocaproyl-valine-citrulline-((paminobenzyl)oxy)carbonyl linker (MC-PV-PABC) by reduction of the interchain disulfide bonds (13–15) .
Experimental
ADC Sample Preparation
For intact analysis of ADCs, BV was systematically desalted to remove any components in the formulation buffer before infusion. Samples were buffer exchanged with 200 mM ammonium acetate buffer, pH 7.0, using Amicon centrifugal filter (cutoff = 10,000 Da) (Merck Millipore). Middle-up characterization of ADCs is based on proteolysis by IdeS (FabriCATOR, Genovis) to obtain two Fc/2 fragments and one F(ab′)2 fragment (16). The primary structure characterization of ADCs by sheathless CE–ESI–MS/MS is performed using an adapted bottom-up proteomic experiment. For this reason, the ADC samples are first subjected to tryptic digestion using an in-solution protocol. Note that to maximize digestion yield, BV samples were first denatured using RapiGest surfactant (Waters). To conserve hydrophobic drug-loaded peptides in solution, 10% (v/v) of acetonitrile is added during the reduction step and 40% (v/v) of isopropanol is added at the end of the protocol. After proteolytic digestion, further sample cleanup, especially with solid-phase extraction (SPE), should be avoided because in most cases it leads to loss of the most hydrophilic peptides. Further cleanup generally is not required for the sheathless CE–ESI–MS/MS experiments. The peptide mixture obtained is then diluted to the desired final concentration (2.2 µM) using 50 mM ammonium acetate (pH = 4.0).
CE–ESI–MS hyphenation
For ADC primary structure characterization, we used a CESI8000 (Sciex Separations) sheathless CE–ESI–MS system. This instrument relies on a sheathless interfacing between the CE capillary and the MS system. No sheath liquid is necessary to maintain the coupling, which guarantees an optimal sensitivity because of the low flow rate (<50 nL/min) generated during the separation and the low internal diameter (30 µm) (17). These characteristics tremendously favor the ionization process and provide enhanced sensitivity and a positive impact on the MS data generated. Using hydrodynamic injection, a volume of 91 nL is injected into the the 92 cm x 30 µm bare fused-silica capillary of the CE system. The background electrolyte (BGE) used to perform the electrophoretic separation was composed of 10% acetic acid. Separations were performed by applying a voltage of -20 kV. For nanoelectrospray ionization (nanoESI) infusion experiments, the CE system was hyphenated to a maxis 4G mass spectrometer (Bruker Daltonics). MS transfer parameters were optimized using the actual sample directly infused via the CE system using a pressure of 5 psi (340 mbar). The CE system was coupled to a 6600 TripleTOF MS system (Sciex) equipped with a hybrid analyzer composed of a quadrupole followed by a time-of-flight (TOF) mass analyzer for analysis of BV digests.
For data treatment, intact protein MS analysis or peptide identification was automated for the most part using conventional search algorithms such as Mascot, Biotools (Bruker Daltonics), or BiopharmaView (Sciex, Redwood Shore), which is dedicated to this type of characterization.
Results and Discussion
Drug–Antibody Ratio Determination
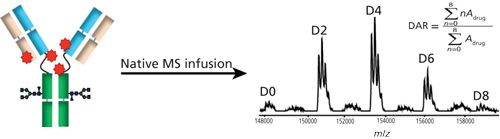
Sheathless CE–MS interface has demonstrated its suitability as a nanoESI emitter to increase sensitivity and limit ion suppression (18). These properties make this system an analytical tool of choice to characterize biologics by direct infusion MS. For the first level of ADCs characterization, the drug load distribution and the average DAR of BV was determined by native MS analysis on the Maxis 4G using nanoESI infusion. Within 5 min infusion at 50 nL/min, MS spectra obtained show a distribution of baseline resolved species corresponding to the intact brentuximab linked with zero to eight payloads (Figure 1) (12). Free light chain (Lc) or heavy chain (Hc) loaded with the drugs, potentially formed during the ionization or the ion transfer inside the mass spectrometer could not be detected. The generated data show that implementation of the sheathless CE–MS interface is compatible with native MS conditions. NanoESI infusion improves the detection of the different species and allows to obtain the drug-load distribution. Using charge state deconvoluted mass spectrum of intact and deglycosylated BV, average DAR were calculated using the relative quantitation from the peak area of each peak and the corresponding number of drug loaded. Average DAR values of 3.8 and 3.9 respectively were determined which are in a good agreement with literature values (19–21). Results demonstrated, using the sheathless CE–MS system as a nanoESI platform, the possibility to perform the analysis of ADCs in native conditions. Therefore confident structural information (number of loaded drugs), species distribution and average DAR could be deducted from a single infusion that required less than 1 µg of intact BV.
Figure 1: Native MS infusion for average DAR determination and drug loaded distribution assessment.
Amino Acid andGlycosylations Characterization
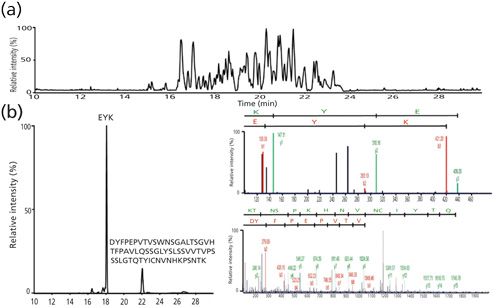
Based on a bottom-up proteomic approach, BV samples were treated using an adapted in solution digestion protocol. In a previous study, our group described the development of a CE–MS/MS method to perform the simultaneous characterization of mAbs with respect to several aspects that define their primary structure, including amino acid sequence, glycosylation, and other types of PTMs (22). However, the presence of hydrophobic drugs linked to the peptide backbone of brentuximab prevented the detection of these drug-loaded peptides because of their precipitation. Based on a previous study (23), we adapted our standard in-solution tryptic digestion with several modifications to make it compatible with the electrophoretic separation. Before trypsin proteolysis, IdeS enzymatic reaction was performed to cleave BV between the two consecutive glycine residues present in the hinge region. Before trypsin digestion, 10% (v/v) acetonitrile was added to the sample, and 40% (v/v) isopropanol was added after digestion. BV digests were analyzed through sheathless CE–MS/MS using a TOF-MS system with an injection volume corresponding to 200 fmol of digested peptides. Peptides were identified based on both mass measurement and MS/MS spectra similarly to bottom-up proteomic experiments. Sheathless CE–MS/MS data from a single injection provided a sequence coverage of 100% for both the Hc and Lc of BV (12). The ability to characterize completely the amino acid sequence is mainly attributed to the properties of CE separation. Indeed, peptides are migrating under the influence of the electrical field applied regardless of their chemical nature, which ensures their successful transfer to the MS system. As a result, it was possible under the same conditions to identify both very small peptides (≤3 amino acids) and a 63-amino-acid peptide having a molecular mass greater than 6.6 kDa (Figure 2).
Figure 2: (a) Base peak electropherogram corresponding to the analysis by CE–ESI-MS of a brentuximab tryptic digest. (b) Extracted ion electrophorogram (EIE) corresponding to the mass-to-charge ratios of [EYK] and [DYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTK]. MS/MS fragmentation spectra for both peaks are shown at right. (Adapted with permission from reference 12.)
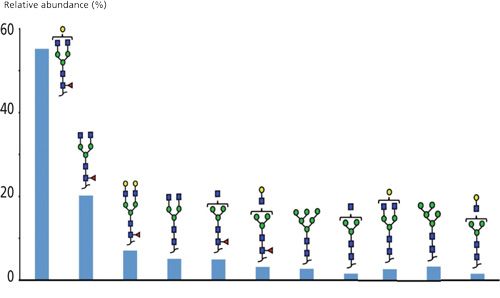
Glycosylation is a factor in mAb safety and pharmacokinetics–pharmacodynamics (PK–PD) and is one of the main sources of heterogeneity for this type of protein (24). Consequently, extensive characterization of glycosylation in regarding to its structure and relative abundance is mandatory. Because it is the combination of a cytotoxic drug and a monoclonal antibody, BV is N-glycosylated on Asn-297 in the heavy chain. The same data set could be used for amino acid sequence identification and determination of the N-glycosylation of BV. The sheathless CE–MS interface demonstrated an excellent ionization efficiency, and therefore glycopeptides could be detected with an outstanding sensitivity. In addition, the MS/MS spectral quality enabled confident determination of the structure of the glycans. Identifications were based on accurate mass measurement in MS1 provided by high-resolution MS and additionally by fragmentation spectra. Results demonstrated the possibility of this analytical approach to obtain simultaneously the structural characterization of 11 glycoforms for BV. The ability to separate glycopeptides with these faint differences is definitely an asset. In the case of comigration, they tend to compete against each other during the ionization process, which may lower the sensitivity of the signal and introduce biases to the relative quantification. Therefore, structural glycan characterization as well as relative quantification could both be completely integrated into the CE–MS characterization workflow using the same set of data (Figure 3).
Figure 3: Glycoform relative abundances determination obtained for brentuximab vendotin using the CE–ESI-MS/MS method in a single analysis.
This work confirms our previous studies on the characterization of mAb primary structure using sheathless CE–MS (22) and demonstrates the possibility of using the approach to characterize the ADC amino acid sequence and glycosylation profile. The modifications made to the sample preparation protocol do not affect the performance of the method on the amino acid sequence characterization compared to results previously published (22). The presence of significant amounts of organic solvents during the reduction and digestion steps of the sample preparation does not prevent the electrophoretic separation or the detection of the tryptic peptides.
Characterization of Drug-Loaded BV Peptide
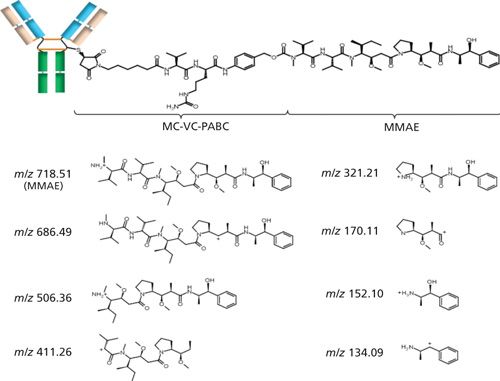
Compared to mAbs, ADCs require an additional level of characterization that focuses on the conjugated drug. When the characterization is performed through peptides after proteolytic digestion, the hydrophobicity caused by the presence of the drug induces precipitation of drug-loaded peptides in aqueous buffers such as ammonium bicarbonate. Indeed, when applying our protocol based on a conventional in-solution trypsin digestion, developed in a previous work for mAbs characterization (22), no detection of drug-loaded peptides was obtained. To prevent precipitation, organic solvents have been added during reduction and digestion steps of the sample preparation. Using these precautions, drug-loaded peptides generated from the tryptic digestion of BV could be maintained in solution and successfully identified by sheathless CEMS. In this case as well, identification of each drug-loaded peptide is based on accurate mass measurement in MS and MS/MS spectra. However, the most abundant product ions observed in MS/MS with low-energy collision-induced dissociation (CID) mainly correspond to the fragmentation of the MMAE drug moiety. In addition to the characteristic in-source fragmentation ion (m/z 718.51) recently reported (25), seven new diagnostic ions corresponding to the activation of the MMAE have been identified and modeled (Figure 4). The presence of several characteristic ions attributed to drug-loaded peptides therefore improves the confidence regarding the localization of the drug on the peptide backbone. As a consequence it was possible to characterize four different drug-loaded peptides corresponding to the Lc peptide [GEC*], the Hc peptide [SC*DK], and the Hc peptide [THTC(*)PPC(*)PAPELLG]-all conjugated with one payload-and the Hc peptide [THTC*PPC*PAPELLG], which is conjugated to two payloads. In each case, cysteine residues present on the peptide backbone are involved in linkage with the MMAE drug.
Figure 4: Schematic structures of brentuximab vedotin and characteristic fragments of MMAE payload. (Adapted with permission from reference 12.)
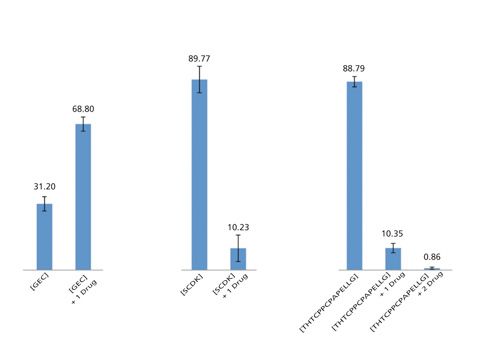
Those results then confirm the characterization of the expected drug-loaded peptides linked to the interchain cysteines of the Hc and Lc (23). Moreover, similar to glycosylation characterization, the electrokinetically driven separation provided by CZE enables the separation of unmodified peptides from their modified counterparts. In this context, the separation of native peptide and the same peptide bearing one or two payloads permits the estimation of the relative abundance of each form and the site by site yield of drug incorporation (Figure 5).
Figure 5: Drug-loaded-peptide relative abundances determination obtained for brentuximab vendotin using the CE–ESI-MS/MS method in a single analysis.
Conclusion
The emergence of biotherapeutic proteins in this demanding regulatory environment creates a challenge for the analytical sciences because of their structural complexity and the potential presence of various types of PTMs. Compared to mAbs, ADCs possess an additional level of complexity due to the cytotoxic drug conjugated on the peptide backbone of the mAb. Starting with an intact ADC native MS study and going through a bottom-up MS approach, we were able to characterize BV from intact mass measurement to the amino acid sequence level. At the intact ADC level, we evaluated a sheathless CE–MS system used as a nanoESI infusion platform for the native MS determination of the average DAR and drug load distribution of BV. Within a single run with a duration of just a few minutes, the technique enabled calculation of DAR values that were in good agreement with previously published values. Using sheathless CE–MS/MS for bottom-up analysis, it is possible from a single injection corresponding to 200 fmol of digested ADCs to characterize the amino acid sequence of the protein systematically with complete sequence coverage. Also, the glycoprofile of the ADCs can be successfully established with the same MS/MS dataset. The glycoprofile includes the structural characterization of expressed glycosylation in addition to the estimate of relative abundance of the various glycoforms. Finally the very same data can also be used to locate the conjugated drug on the peptide backbone and estimate the yield of the coupling reaction independently for each conjugation site. MS/MS spectra corresponding to drug-loaded peptides allowed for the first time the identification of several diagnostic ions strictly attributed to the fragmentation of the drug. The presence of diagnostic ions prevents ambiguity concerning the characterization of drug-loaded peptides.
The versatility provided by the sheathless CE–MS/MS method results from the combination of the intrinsic characteristics of the CE–MS interfacing and the implementation of an electrokinetic separation. Indeed the design of the sheathless CE–MS interface supported separation conditions that proved to be favorable to the ionization process and provided successful transfer of the produced ions to the MS. Therefore it is responsible for the exquisite sensitivity achieved, particularly for glycosylation and PTM characterization. Also, the CE separation mechanism demonstrated the ability to separate peptides having minor differences, which limits the introduction of bias during the ionization process and improves the relative quantification. The outstanding level of performance achieved in a minimum number of experiments positions this sheathless CE–MS analytical workflow as a powerful tool even for R&D support activities where information about the studied ADC is scarce.
Acknowledgments
Authors would like to thank Sciex separations Inc. for lending a CESI8000 system and a 5600 TripleTOF mass spectrometer. Authors would like to thank Bruker Daltonics for lending a Maxis 4G system. The authors thank Jim Thorn, Steven Lock, and Dominique Mellon from Sciex separations Inc. and Yann Hebert, Pierre-Olivier Schmit, and Jean-Michel Billmann from Bruker Daltonics for their support. The authors would like also to express their gratitude to Dr. M. Biacchi (LSMIS, Strasbourg, France), P. Hammann, and J. Chicher (Institut de Biologie Moléculaire et Cellulaire, Strasbourg, France), Dr. E. Wagner-Rousset, , MC. Janin-Bussat, and O. Colas (Centre d’Immunologie Pierre Fabre, St Julien en Genevois, France) for helpful discussions. This work was supported by the CNRS (UMR 7140), the University of Strasbourg, and a doctoral fellowship from the General Council of Mayotte to NS.
References
- S. Ornes, Proc. Natl. Acad. Sci. U. S. A.110, 13695–13695 (2013).
- A. Beck, J.M. Reichert, mAbs6, 15–17 (2014).
- R.S. Zolot, S. Basu, and R.P. Million, Nat. Rev. Drug Discovery12, 259–260 (2013).
- U. Iyer and V.J. Kadambi, J. Pharmacol. Toxicol. Methods64, 207–212 (2011).
- S. Krapp, Y. Mimura, R. Jefferis, R. Huber, and P. Sondermann, J. Mol. Biol.325, 979–989 (2003).
- A. Beck, G. Terral, F. Debaene, E. Wagner-Rousset, J. Marcoux, M.-C. Janin-Bussat, O. Colas, A. Van Dorsselaer, S. Cianférani, Expert Review of Proteomics2, 1–27 (2015).
- B. Yan, J. Valliere-Douglass, L. Brady, S. Steen, M. Han, D. Pace, S. Elliott, Z. Yates, Y. Han, A. Balland, W. Wang, and D. Pettit, J. Chromatogr. A1164, 153–161 (2007).
- R. Gahoual, A. Beck, Y.-N. François, and E. Leize-Wagner, J. Mass Spectrom.51, 150–158 (2016).
- S. Fekete, D. Guillarme, P. Sandra, and K. Sandra, Anal. Chem.88, 480–507 (2016).
- R. Gahoual, A. Burr, J.M. Busnel, L. Kuhn, P. Hammann, A. Beck, Y.N. Franccois, and E. Leize-Wagner, mAbs5, 479–490 (2013).
- R. Gahoual, M. Biacchi, J. Chicher, L. Kuhn, P. Hammann, A. Beck, E. Leize-Wagner, and Y.N. Francois, mAbs6, 1464–1473 (2014).
- N. Said, R. Gahoual, L. Kuhn, A. Beck, Y.-N. François, and E. Leize-Wagner, Anal. Chim. Acta918, 50–59 (2016).
- A. Younes, N.L. Bartlett, J.P. Leonard, D.A. Kennedy, C.M. Lynch, E.L. Sievers, and A. Forero-Torres, N. Engl. J. Med.363, 1812–1821 (2010).
- J.A. Francisco, C.G. Cerveny, D.L. Meyer, B.J. Mixan, K. Klussman, D.F. Chace, S.X. Rejniak, K.A. Gordon, R. DeBlanc, B.E. Toki, C.L. Law, S.O. Doronina, C.B. Siegall, P.D. Senter, and A.F. Wahl, Blood102, 1458–1465 (2003).
- E. Oflazoglu, K.M. Kissler, E.L. Sievers, I.S. Grewal, and H.P. Gerber, Br. J. Haematol.142, 69–73 (2008).
- Y.-N. François, M. Biacchi, N. Said, C. Renard, A. Beck, R. Gahoual, and E. Leize-Wagner, Anal. Chim. Acta908, 168–176 (2016).
- .M. Busnel, B. Schoenmaker, R. Ramautar, A. Carrasco-Pancorbo, C. Ratnayake, J.S. Feitelson, J.D. Chapman, A.M. Deelder, and O.A. Mayboroda, Anal. Chem.82, 9476–9483 (2010).
- R. Gahoual, J.M. Busnel, P. Wolff, Y.N. Francois, and E. Leize-Wagner, Anal. Bioanal. Chem.406, 1029–1038 (2014).
- F. Debaene, A. Boeuf, E. Wagner-Rousset, O. Colas, D. Ayoub, N. Corvaia, A. Van Dorsselaer, A. Beck, and S. Cianferani, Anal. Chem.86, 10674–10683 (2014).
- J.F. Valliere-Douglass, W.A. McFee, and O. Salas-Solano, Anal. Chem.84, 2843–2849 (2012).
- J. Chen, S. Yin, Y. Wu, and J. Ouyang, Anal. Chem.85, 1699–1704 (2013).
- R. Gahoual, J.-M. Busnel, A. Beck, Y.-N. François, and E. Leize-Wagner, Anal. Chem.86, 9074–9081 (2014).
- M.-C. Janin-Bussat, M. Dillenbourg, N. Corvaia, and A. Beck, J. Chromatogr., B 981, 9–13 (2015).
- A. Beck, J.M. Reichert, mAbs4, 419–425 (2012).
- J.R. Junutula, H. Raab, S. Clark, S. Bhakta, D.D. Leipold, S. Weir, Y. Chen, M. Simpson, S.P. Tsai, M.S. Dennis, Y. Lu, Y.G. Meng, C. Ng, J. Yang, C.C. Lee, E. Duenas, J. Gorrell, V. Katta, A. Kim, K. McDorman, K. Flagella, R. Venook, S. Ross, S.D. Spencer, W. Lee Wong, H.B. Lowman, R. Vandlen, M.X. Sliwkowski, R.H. Scheller, P. Polakis, and W. Mallet, Nat. Biotechnol. 26, 925–932 (2008).
Nassur Said, Emmanuelle Leize-Wagner, Jérémie Giorgetti, and Yannis-Nicolas François are from the Laboratory of Mass Spectrometry of Interactions and Systems (LSMIS), UMR 7140 (Unistra-CNRS), University of Strasbourg, Strasbourg, France. Lauriane Kuhn is from the Strasbourg-Esplanade Proteomic Platform, Institute of Molecular and Cellular Biology, at the Univeristy of Strasbourg, Strasbourg, France. Alain Beck is from the Pierre Fabre Immunology Center in Saint-Julien-en-Genevois, France. Rabah Gahoual is from the Unit of Chemical and Biological Technologies for Health at University of Paris Descartes in Paris, France.

Newsletter
Get essential updates on the latest spectroscopy technologies, regulatory standards, and best practices—subscribe today to Spectroscopy.



