- Spectroscopy-11-01-2019
- Volume 34
- Issue 11
Quantum Cascade Laser Infrared Microscopy and 2D IR Correlation Spectroscopy Improves Crystallization Screening of a Protein Complex
Well-diffracting crystals are essential for X-ray diffraction of crystallized protein for structural determination. A quantum cascade laser (QCL) infrared microscope is used to determine protein aggregation, distinct from self-association, for the success of the crystallization effort.
X-ray diffraction of crystallized protein continues to be the preferred means of obtaining high-resolution structures of proteins and their complexes. These structures are crucial for drug design, but the screening and optimization of the conditions that produce well-diffracting crystals represent a bottleneck in the structure determination process. In this study, a quantum cascade laser (QCL) infrared microscope was used to determine protein aggregation, distinct from self-association, which is crucial to the success of any crystallization effort. Hyperspectral images of an aliquot from a vapor diffusion hanging drop crystallization screen were acquired over a small temperature range (30–38 ºC), at intervals of 2 ºC. QCL infrared (IR) spectral data were subjected to 2D IR correlation analysis to describe Homo sapiens (Hs) centrin 2(E32A)-Sfi1p21 complex (1:1.5 molar ratio) and the selective aggregation of the target peptide. To our knowledge, this is the first time such a level of molecular understanding has been achieved.
Protein crystallization with X-ray diffraction (XRD) is the preferred means of obtaining high-resolution structures of proteins and their complexes. These structures are crucial for drug design, yet methodological barriers to their determination still exist (1−4). Efforts to make the structure determination process more efficient include data collection methods, such as the synchroton beam lines (5), detection via sensitive focal plane arrays, and improved cryogenic and mounting procedures for crystals. However, one key step in successful high-throughput X-ray crystallography continues to be a bottleneck; namely, the screening and optimization of the conditions required to produce well-diffracting crystals. This process includes the optimization of multiple variables, including, but not limited to, precipitant, pH, temperature, protein sequence, and concentration (4). New techniques are needed if we hope to increase the rate of success of this step, which is currently ~18%.
There are three stages in the crystallization process: nucleation, crystal growth (which is governed by diffusion), and cessation of crystal growth. Crystal growth requires specific interactions between individual protein molecules that lead to an organized crystal lattice. If the crystallizability of the protein of interest is dependent on the type of protein aggregate generated during the crystallization process, then our proposed method should allow for this assessment. Conceptually, the aggregate should be one of self-association, in which chemical associations driven by the weak interactions between proteins or the precipitating agent are critical to maintaining the protein’s conformational stability. Self-association in principle would account for the generation of a crystal lattice, while excluding solvent molecules from the immediate hydration shell of the protein and increasing the influence of the surface charge of the protein to account for the crystallization event. The evaluation of the crystallization process of proteins and protein complexes using this direct and highly informative approach would allow for further understanding of the process of crystallization, and should be explored even further during crystallization optimization as part of the screening process. It may also be possible to evaluate the precipitant’s role in changing the dynamics of the protein’s interaction with its aqueous environment, causing a synergistic effect that will lead to the successful crystallization of the protein or the protein complex.
Here, we have employed quantum cascade laser infrared microscopy (QCL IRM) for the determination of protein aggregation, distinct from self-association, under various crystallization conditions. The platform includes a QCL infrared (IR) microscope with enhanced signal-to-noise ratio, sampling accessories, cells under thermal control, and software for QCL IR spectral analysis and assessment of crystallizability. Hyperspectral images (HSI) within the mid-IR spectral region of 1800–1000 cm-1 provide data distinguishing an aggregate from a crystal, as well as a rough determination of the crystal or aggregate size. The QCL IR spectral data are subjected to 2D IR correlation spectroscopy analysis (6,7) to determine the extent and molecular mechanism of aggregation and crystallization under thermal stress.
A high-resolution structure for a yeast homologue of centrin known as CDC31 in the form of CDC31-Sfi1p complex exists as PDB ID: 2DOQ (Figure 1) (8), and our group has extensively studied Hscentrin isoforms using multiple biophysical techniques, including differential scanning calorimetry and circular dichroism, Fourier transform infrared (FT-IR), and two-dimensional (2D) IR correlation spectroscopies (9). The current study involved a complex between a centrin variant (Hscentrin 2 [E32A]) and its target peptide, Sfi1p21, which is presented as proof-of-concept. In this work, 2D IR correlation spectroscopy provided molecular insight into which of the components was aggregated and how the initial interaction of Sfi1p21 with the centrin variant led to the selective aggregation of the peptide. To our knowledge, this is the first time that such a level of molecular understanding has been achieved for a crystal screen. The results obtained demonstrate the breakthrough capabilities of the QCL IR microscope along with 2D IR correlation analysis for the evaluation of the crystallizability of a protein or protein complex.
Experimental
Crystal Screen Setup
We performed a vapor diffusion hanging drop crystal screen containing 2 µL of the Hscentrin 2 (E32A) variant and HsSfi1p21 synthetic peptide (1:1.5 molar ratio) for a 5 mM total protein concentration and 2 µL of the index screen from Hampton Research were placed in a vapor diffusion setup tray from the Easy Xtal Tool from Qiagen Sciences. The visual evaluation of the screen was done using a Nikon SMZU microscope for selection of potentially diffracting crystals. We identified one condition to contain both amorphous and microcrystals: condition #31 of the index screen. This crystallization solution contained 0.1 M Tris at pH 8.5, 0.5% (w/v) polyethylene glycol monomethyl ether 5000, and 0.8 M potassium sodium tartrate tetrahydrate.
A 1 µL aliquot from a selected hanging drop, identified to contain amorphous or microcrystals, was drawn from the hanging drop, placed in a predefined well of a custom milled calcium fluoride slide, and covered with a fully polished crystal to make up the slide cell. The slide cell was then placed in a heated accessory with accurate thermal control, and HSIs were acquired within the temperature range of 30–38 ºC with 2 ºC intervals and 4 min equilibration periods using the ProteinMentor, a QCL IR microscope from Protein Dynamic Solutions, Inc.
Hyperspectral Image (HSI) Acquisition
The QCL infrared microscope acquires the HSIs, allowing for a linear response microbolometer focal plane array (480 x 480 pixels) detector to be used. A low magnification objective (4x), with a numerical aperture (NA) of 0.3 and a 2 x 2 mm2 field of view, provides a pixel size of 4.25 x 4.25 µm2. The HSIs are composed of 223,000 QCL IR spectra and were collected at 4 cm-1 resolution within the spectral region of 1750−1500 cm-1. To prevent coherence effects due to QCL IR fluctuations, the background was collected at each temperature once thermal equilibrium (4 min) was achieved.
QCL IRM spectral overlay and 2D IR correlation analysis for this sample was performed using the Kinetics program of MATLAB, which was generously provided by Dr. Erik Goormaghtigh from the Free University of Brussels, Belgium. However, we have developed fully automated Correlation Dynamics software to analyze an array of samples.
Results and Discussion
Size of the Aggregate
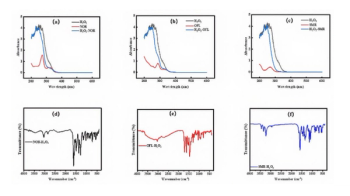
HSIs for the Hscentrin 2 (E32A)-Sfi1p21 sample at five different temperatures (30, 32, 34, 36, and 38 ºC) are shown in Figure 2. Initially, microcrystals were observed in a circular arrangement, due to the dispensing of the pipette, which allows for the amorphous or microcrystals to flow from the tip to the well during loading (Figure 2). A micrometer scale was used to determine the size of the aggregates observed in the HSI; the size of the aggregate was determined to be 30 µm x 40 µm at 30 ºC. Furthermore, as the temperature was increased, the aggregate continued to grow, reaching 300 µm x 400 µm in size at 38 ºC. The corresponding QCL IR spectra (Figure 3a) were indicative of the presence of aggregate, thus confirming the HSIs (Figure 2). Specifically, the full width at half height of the amide I band and the presence of a shoulder at 1613 cm-1 are consistent with the presence of aggregates. While 2D IR correlation analysis yielded the typical synchronous plot observed for a protein aggregation process during thermal perturbation (as indicated by the prominent auto peak 1613 cm-1 present in the synchronous plot), the asynchronous plot suggested an unfolding event during the aggregation process, as indicated by the cross peaks located within 1613 cm-1.
Band Assignments
2D IR correlation spectroscopy was performed to improve our understanding of the backbone dynamics and side chain interactions involved in the aggregation process within a crystallization condition. Band assignments were made using the amino acid sequence of Hscentrin 2, obtained from the National Center for Biotechnology Information database (NCBI accession number: EAW72900). The band assignments are summarized in Table I. The vibrational modes associated with Hscentrin 2 (E32A) include the p-aromatic overtones for phenylalanine residues (1702.0 and 1717.5 cm-1) and calcium binding site loops (1682.6 and 1672.0 cm-1). EF-hand proteins also have hinge loops, which we have assigned to 1662.5 cm-1. We also assigned the helical component (1652.5 cm-1), and the short β-sheet segments (1632.0 cm-1). Finally, the aspartate carboxylate stretching side chain vibrational mode ν (COO-), mainly located within the calcium binding sites within centrin, was assigned to 1577.0 cm-1. Shared vibrational modes were the glutamate carboxylate side chain mode ν(COO-) (1550.3 cm-1), found in both centrin and the Sfi1p21 target peptide. Exclusive to Sfi1p21 were two vibrational modes due to aggregation (1613.0 cm-1) and the two sets of contiguous histidine residues with the stretching vibrational mode ν(C=C) (1597.2 cm-1). These band assignments are consistent with the secondary structure information available in the high-resolution crystal structure of the complex and with previous work from our laboratory (8,9). Also, the side chain modes and their molar extinction coefficients have been determined (10) and reviewed (11,12).
2D IR Correlation Analysis
The protein sample is in an aqueous environment and the molar extinction coefficient of pure H2O is high at 55.5 M-1 cm-1, yet, like any protein-containing sample, it has effectively diluted the contribution of H2O in the overall absorbance spectrum. Also important is the decreased pathlength, allowing for the management of samples that exhibit high absorptivity. Finally, the QCL IRM transmission absorbance spectra have an enhanced signal-to-noise ratio allowing for the difference spectra approach to be used, thus overcoming the common deterrents of using IR spectroscopy for aqueous samples. The QCL IR spectrum for the region of interest at the initial temperature (30 ºC) was subtracted from all subsequent spectra, thereby generating the difference spectra. It is to this data set that the 2D IR correlation function is applied to generate the synchronous and asynchronous plots.
Difference spectra used in 2D IR correlation spectroscopy are defined as:
where Ä(υj) is the initial spectrum of the data set to generate the covariance spectra. Synchronous 2D correlation intensities of the covariance spectral data are defined by:
where the resulting correlation intensity Φ(ν1,ν2) as a function of two independent wavenumber axes, ν1 and ν2, is the synchronous plot.
Asynchronous 2D correlation intensities of the covariance spectral data are defined by:
where the term Nij is the element of the so-called Hilbert-Noda transformation matrix, given by:
These 2D IR correlation plots provide enhanced resolution of the peak intensity changes and positions within the spectral region of interest that were due to the temperature increase. Specifically, the synchronous plot was used to establish relationships between peaks that change in the same phase with one another (Figure 3b). This plot has peaks on the diagonal known as auto peaks, which provide the main peak intensity changes. In addition, peaks off the diagonal provide the relationship between the auto peaks and are known as cross peaks. In this case, as shown in Figure 3b, the only intensity change was at 1613 cm-1, and was due to the aggregation process. The synchronous plot did not contain any negative peaks, thus simplifying the analysis even further. The asynchronous plot (Figure 3c) establishes the relationship between peaks that are changing out-of-phase from one another. In general, there are no peaks on the diagonal and only cross peaks are observed. This plot provides enhanced resolution, and is used to determine the sequential order of molecular events that describe the aggregation process during the temperature perturbation. The order of molecular events was determined by following Noda’s rules (6,7). Keeping in mind that asynchronous plots are symmetrical in nature, and by convention we always refer to the top triangle for analysis, we apply the rules as follows:
I. If asynchronous cross peak ν2 is positive, then ν2 is perturbed prior to ν1 (ν2 → ν1).
II. If asynchronous cross peak ν2 is negative, then ν2 is perturbed after to ν1 (ν2 ← ν1).
III. If the corresponding synchronous cross peak is positive, then the order of the events is established using the asynchronous plot (rules I and II).
IV. However, if the corresponding synchronous cross peak is negative and the asynchronous cross peak is positive, then the order is reversed.
The order of events can be established for each peak observed in the ν2 axis (Figure 3c). In this specific case, the only two rules that apply are rules I and II. Our group has successfully applied 2D IR correlation spectroscopy to the study of numerous proteins and peptides, including a comparative analysis of related human centrins (9).
Description of the Molecular Aggregation Process
Figure 4 gives a schematic representation of the molecular events that lead to the initial interaction of Sfi1p21 with Hscentrin 2 (E32A) and the selective aggregation of the target peptide, based on the features of the asynchronous plot. Initially, the glutamate residues (1550.3 cm-1), which could be attributed to either Hscentrin 2 (E32A) or the Sfi1p21 target peptide, were perturbed, followed by the aspartates (1577.0 cm- 1), which are exclusive to centrin and are located within the calcium binding sites. Next, the short β-sheets (1632.0 cm-1) that are located close to the calcium binding sites were perturbed, presumably due to the initial interaction with the target peptide. At this point, the optimum orientation was not achieved between the Hscentrin 2 (E32A) and Sfi1p21, leading to the selective aggregation (1613 cm-1) of the target peptide via its contiguous histidine residues (1597.2 cm-1), located both in the middle of the sequence (H13-H15, ε = 210 M-1 cm-1) and near the C-terminal end (H29-H30, ε = 140 M-1cm-1). These histidine vibrational modes are exclusive to Sfi1p21, and the histidine pairs are presumably involved in the aggregation process as two separate but simultaneous molecular events during the thermal stress. We theorize that the Hscentrin 2 (E32A) variant may predispose the initial interaction with Sfi1p21 to the EF-hand motif located in the N-terminal end instead of the EF-hand motif located within the C-terminal end, preventing the optimum orientation that would normally lead to Sfi1p21 adopting a helical conformation. The helical region of Hscentrin (E32A) (1652.5 cm-1) was then perturbed, followed by opening of the EF-hand motif, which perturbs the hinge loop (1662.5 cm-1) and associated calcium binding site loops (1672.0 and 1682.6 cm-1, which represent the apo- and holoforms of the calcium binding sites, respectively). The opening of the EF-hand exposes the phenylalanine residues (1715 and 1702 cm-1, p-aromatic overtones) to their aqueous environment, leading to an unstable centrin variant-target peptide interaction which resulted in the selective aggregation of Sfi1p21.
Conclusion
The QCL IRM platform, combined with 2D IR correlation spectroscopy, has proven useful for obtaining a molecular description of the aggregation distinct from the desired self-association of a protein-peptide complex during crystal screening. A successful crystal screen would include the self-association of proteins or the association observed in protein complexes, which would lead to nucleation and crystal growth. 2D IR correlation spectroscopy would be capable of distinguishing between the two types of processes, while providing an unprecedented level of molecular detail. This approach may provide valuable information leading to increased success rates for the crystallization of protein complexes by identifying the variables that lead to the optimum chemical and physical properties associated with well-diffracting crystals.
Author Contributions
Crystallization screens were performed and HSIs acquired by Sherly Nieves. The 2D IR spectral data analysis was performed by Belinda Pastrana. Figures were generated by Sherly Nieves.
Acknowledgments
The authors would like acknowledge Dr. Melissa Stauffer (Scientific Editing Solutions, Walworth, WI) for editing the manuscript.
Funding
The work presented herein was made possible by support from the National Science Foundation (SBIR PII Award 1632420 [BP]).
Disclosures
Belinda Pastrana is the CEO of Protein Dynamic Solutions, Inc. and a Professor in the Department of Chemistry at the University of Puerto Rico-Mayagüez, Mayagüez, PR. Sherly Nieves is a scientist at Protein Dynamic Solutions, Inc.
References
- I.D. Hoffman, Methods Mol. Biol.841, 67–91 (2012).
- I. Russo Krauss, A. Merlino, A. Vergara, and F. Sica, Int. J. Mol. Sci. 14, 11643–11691 (2013).
- C. Sauter, B. Lorber, A. McPherson, and R. Giege, in International Tables for Crystallography. Volume F: Crystallography of Biological Macromolecules, E. Arnold, D.M. Himmel, and M.G. Rossman, Eds. (Wiley, Chichester, New Hampshire, 2nd ed., 2012), pp. 99–121.
- Y. Lin, Expert Opin. Drug Discovery 13, 691–695 (2018).
- R.A. Judge, S. Takahashi, K.L. Longenecker, E.H. Fry, C. Abad-Aapatero, and M.L. Chiu, Cryst. Growth Des.9, 3463–3469 (2009).
- I. Noda, Vibrat. Spectrosc.36, 143–165 (2004).
- I. Noda, J. Mol. Struct.1069, 23–49 (2014).
- S. Li, A.M. Sandercock, P. Conduit, C.V. Robinson, R.L. Williams, and J.V. Kilmartin, J. Cell. Biol.173, 867–877 (2006).
- B. Pastrana-Rios, J. De Orbeta, V. Meza, M. Reyes, D. Narváez, A.M. Gómez, A. Rodríguez Nassif, R. Almodovar, A. Díaz Casas, J. Robles, A.M. Ortíz, L. Irizarry, M. Campbell, and M. Colón, Biochemistry 52, 1236–1248 (2013).
- S.Y. Venyaminov and N.N. Kalnin, Biopolymers 30, 1243–1257 (1990).
- J. Bandekar. Biochim Biophys Acta,1120, 123–143 (1992).
- A. Barth. Biochim. Biophys. Acta, 1767, 1073–1101 (2007).
Sherly Nieves and Belinda Pastrana are with Protein Dynamic Solutions, Inc., in Wakefield, Massachusetts. Direct correspondence to:
Articles in this issue
over 6 years ago
Data Quality and Data Integrity Are the Same, Right? Wrong!over 6 years ago
Vol 34 No 11 Spectroscopy November 2019 Regular Issue PDFover 6 years ago
Organic Nitrogen Compounds VI: Introduction to AmidesNewsletter
Get essential updates on the latest spectroscopy technologies, regulatory standards, and best practices—subscribe today to Spectroscopy.

![Figure 3: Plots of lg[(F0-F)/F] vs. lg[Q] of ZNF191(243-368) by DNA.](https://cdn.sanity.io/images/0vv8moc6/spectroscopy/a1aa032a5c8b165ac1a84e997ece7c4311d5322d-620x432.png?w=350&fit=crop&auto=format)

